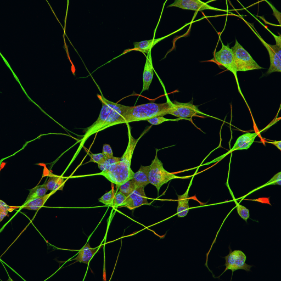
Humans are experiencing stress at higher levels than ever before. In the developed world, this appears to be a product of our way of life, where we are busier than ever yet more isolated from our natural environments and communities. In other places, war, conflict, climate change and environmental disasters are displacing people from their homes and countries at unprecedented rates.
Stress is not always bad for us. It’s what gets us out of bed and gives us laser focus. Yet stress that is stronger than an individual’s ability to adapt and cope is one of the leading risk factors for developing severe mental illnesses including depression, bipolar disorder, schizophrenia, post-traumatic stress disorder, and anxiety. In fact, the World Health Organisation predicts that by 2030, one third of all disease burden in the world will be caused by stress.
We are therefore at a crossroad. We urgently need an improved understanding of the detailed and widespread effects of stress on human biology, so that we can identify people who are vulnerable to the effects of stress and improve their resilience. To do this, we must first understand what are the biological effects of stress and how does stress raise risk to mental illness.
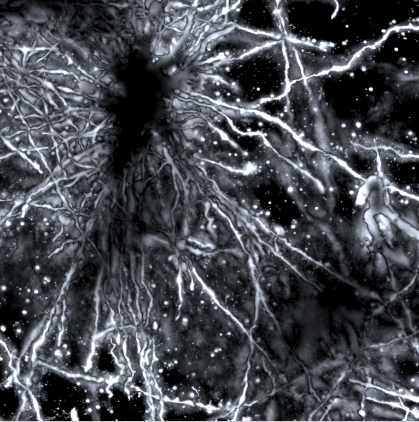
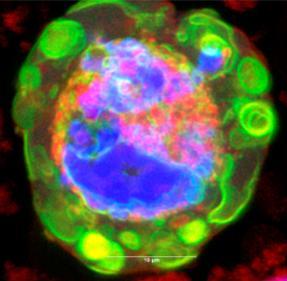
An inverse Colgi-cox stained image of the human brain cortex by Dominical Kaul
Our goal
The Matosin Lab broadly aims to understand how stress contributes to the development of mental illness. The lab has two main streams:
- In the first stream, we aim to understand what happens to the cells and molecules in the human brain after stress exposure or in mental illness. To do this, we study human brains donated to science by people who used to live with a mental illness and/or had very stressful lives. Tiny slivers of brain or pieces no larger than the size of a pea are used to pinpoint differences in the shapes, numbers, orientation and connections of brain cells, as well as what is happening inside them from the level of the gene to the protein. This research provides the fundamental knowledge needed to develop new treatments and interventions.
- In the second stream, we aim to understand what are the long-term and sustained effects of stress on the human body, and then to build a framework for identifying people who are at risk to mental illness and ways to improve their resilience. Our group is also interested in how the effects of stress and trauma can be transmitted from parent to offspring, therefore having transgenerational impact. To address these questions, we study biological samples – including saliva, mouth swabs, blood, and breast milk – and psychological data from people and communities who have been heavily stress exposed. By studying human tissues and fluids that are easily accessible and minimally invasive to collect, this research provides the possibility to develop ways to (a) screen for people at risk to the detrimental effects of stress, (b) identify who could benefit from specific treatments and interventions, and (c) design those treatment and interventions.
Our values in and out of the lab
Scientific excellence, impactful research, collaboration, openness and authenticity, training and connecting the next generation of scientists with world leaders, enthusiasm, creating an environment that is positive and encourages teamwork and generosity.
View Dr Natalie Matosin's Scholars page
Contact nmatosin@uow.edu.au for more information.